„G-kvadruplex” változatai közötti eltérés
Új oldal, tartalma: „{{építés alatt}} Fájl:G-quadruplex.svg|bélyegkép|400px|Egy G-kvadruplex szerkezete. Baloldalt G-tetrád, jobboldalt intramolekuláris G4-komplex<ref>{{cite journal|author=J. A. Capra, K. Paeschke, M. Singh, V. A. Zakian|title=G-quadruplex DNA sequences are evolutionarily conserved and associated with distinct genomic features in Saccharomyces cerevisiae | journal = PLOS Computational Biology | volume = 6 | issue = 7 | pages = e1000861 | date = July 2010 |…” Címkék: & jel tataroz vagy építés alatt sablon kihelyezve |
(Nincs különbség)
|
A lap 2023. október 26., 11:59-kori változata
| Ez a szócikk/szakasz most épül, még dolgoznak az első verzión! |
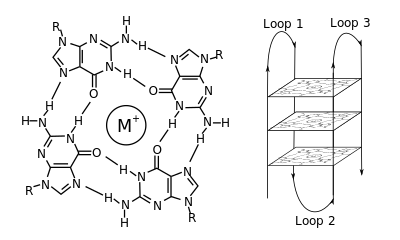
A G-kvadruplex másodlagos szerkezetek (G4) guaninban gazdag szekvenciák által jönnek létre.[2] Ezek helikálisak és 1,[3] 2[4] vagy 4 szálas guanintetrádokból állnak.[5] Az egymolekulás változatok gyakoriak a kromoszómák végén lévő telomereken és számos gén transzkripciószabályzó részein mikrobákban[6][7] és gerincesekben,[8][7] beleértve humán onkogéneket.[9] 4 guanin Hoogsteen-hidrogénkötést alkotva négyzetes szerkezetű guanintetrádot alkothat, és 2 vagy több guanintetrád (folyamatos G-sorozatokból) egymáson G-kvadruplexet alkot.
A G-kvadruplexek elhelyezkedése és kötései nem véletlenszerűek, és különös funkcióik vannak. A kvadruplexet egy tetrádpárok közt középre kerülő kation, különösen a kálium jelenléte stabilizálja.[3] Alkothatja DNS, RNS, ZNS vagy PNS, és lehet molekulán belüli, két- vagy négymolekulás.[10] A szálak vagy azok részeinek ieányától függően a szerkezetek lehetnek párhuzamosak vagy antipárhuzamosak. A G-kvadruplexek DNS- vagy RNS-szekvenciamotívumokból előrejelezhetők,[11][12] de valódi szerkezetük változhat motívumon belül és motívumok közt, amelyekből genomonként több mint 100 000 is lehet. Alapvető genetikai folyamatokban való részvételük miatt kutatják a telomerek, a génszabályzás és a funkciós genomika területén.[13][14]
Története
A nagy guaninasszociációs szerkezeteket 1962-ben azonosították guaninasszociált gélszerű anyagokon keresztül.[15] E kutatás határozta meg a 4 szálú DNS-szerkezetek magas guaninasszociációval való összefüggését, melyeket 1987-ben eukarióta telomerekben fedeztek fel.[16] A G-kvadruplex felfedezésének fontosságát a következő idézet mutatta meg:
| „ | Ha in vitro keletkeznek G-kvadruplexek, a természet alkotott egy módot azok in vivo használatára” | ” |
| – Aaron Klug, az 1982-es kémiai Nobel-díj nyertese | ||
A G-kvadruplexek in vivo funkciójának kutatása a nagyméretű genomszintű analízis hatására megnőtt, melyben a potenciális G-kvadruplex-alkotó (pG4) szekvenciák nagy gyakoriságát mutatták ki humán, csimpánz- egér- és patkánypromoterekben, melyeket a 2007. áprilisi első nemzetközi G-kvadruplex-találkozón mutattak be a kentuckyi Louisville-ben.[7] 2006-ban néhány bakteriális promoterben is kimutattak G-kvadruplexeket, G-kvadruplex-mediált génszabályzást feltételezve.[6] A G-kvadruplexek in vivo gyakorisága révén e szerkezetek biológiailag fontosak az onkogének promotereire és a telomerekre való hatásuk révén. Jelenleg a kutatás a G-kvadruplexek funkcióinak azonosítására bizonyos onkogéneknél, valamint a G-kvadruplexeken alapuló hatékony kezelések felfedezésére irányul. A G-kvadruplexek korai bizonyítéka sejtekből való izolációjuk volt,[17] később annak megfigyelésével is igazolták, hogy egyes DNS-helikázok ilyen szerkezetekhez közel vannak nagy mennyiségben.[18]

Topológia
A tetrádképzésben részt vevő nukleinsavak gatározzák meg a kvadruplex szerkezetét. A rövid, egyetlen 3 vagy több guainból álló sorozatból álló szekvenciák 4 önálló szálat igényelnek a kvadruplexhez, vagyis az ilyen kvadruplex tetramolekulás. A G4 DNS eredetileg az ilyen szerkezetekre utalt, melyek a meiózisban lehetnek fontosak.[5] Azonban a molekuláris biológia modern szóhasznlatában a G4 bármilyen molekularitású kvadruplexet jelenthet. A hosszabb, két folytonos 3 vagy több guaninból álló sorozatot tartalmazó kvadruplexekhez 2 ilyen szekvencia is elég. Ezek a kvadruplexek így bimolekulárisak. Végül a 4 különböző guaninsorozatot tartalmazó szerkezetek önmaguktól képesek stabil kvadruplexet alkotni, így intramolekulárisak.[19]
A guaninbázisok elrendezésétől függően egy kvadruplex számos alakot felvehet különböző konfigurációkkal.[20] Ha minden DNS-szál azonos irányba halad, a kvadruplex párhuzamos. Intramolekuláris kvadruplexek esetén ekkor minden körnek propellernek kell lennie, melyek a kvadruplex oldalain vannak. Ha egy vagy több guaninsorozat 5’-3’ iránya a többivel ellentétes, a kvadruplex antipárhuzamos. Ekkor a guaninsorozatokat egymáshoz csatlakoztató körök az intramolekulás kvadruplexekben vagy átlósak (ellentétes guaninsorozatokat csatlakoztatók), vagy laterálisak (oldal menti), így két szomszédos guaninsorozatot csatlakoztatnak.
A kétszálú DNS-ből alkotott G-kvadruplexekben lehetségesek szálközi topológiák is.[21][22] E kvadruplexek mindkét DNS-szálból tartalmaznak guanint.
Szerkezete és funkciója a genomban
A humán genom szekvenálása után számos kvadruplexalkotásra képes guaningazdag szekvenciát találtak.[23] A sejt típusától és sejtciklusban való állapotától függően a közvetítő tényezők, például a kromatin DNS-kötő fehérjéi és más környezeti tényezők befolyásolhatják a kvadruplexkeletkezést. Például a molekulazsúfoltságg kvantitatív termodinamikai elemzése alapján az antipárhuzamos G-kvadruplexet a molekulazsúfoltság stabilizálja.[24] Ez feltehetően a DNS-hidratáció megváltozása és ennek Hoogsteen-bázispárkötésre gyakorolt hatása okozza.[25] Ezek gyakran kromoszómák végén fordulnak elő. Továbbá a G-kvadruplex-képzés valószínűsége transzkripció során RNS-ben hajtű- vagy G-kvadruplex-szekvenciák létrehozásához a hajtűlétrehozó szekvencia helyétől függ.[26]
Mivel a javító enzimek a lineáris kromoszómák végeit sérült DNS-ként észlelnék, és ezeket káros hatásként dolgoznák fel, egyértelmű jelzés és szigorú szabályzás kell a kromoszómák végein. A telomerek lehetővé teszik e jelzést. Ezek guaninban gazdagok, hajlamosak G-kvadruplex-képzésre, gyakran a kromoszómák végén vannak, és segítik a genom integritásának megőrzését a végek instabilitástól való védelmével.
E telomereket hosszú CCCTAA:TTAGGG ismétlődések jellemzik. Az ismétlődéek 3'-kiemelkedéssel érnek véget 10-50 egyszálú TTAGGG ismétlődéssel. A heterodimer-komplex ribonukleoprotein telomeráz a DNS-szálak 3'-végéhez TTAGGG sorozatokat ad. A G-gazdag meghosszabbítás képes másodlagos szerkezetek, például G-kvadruplexek létrehozására, ha hosszabb 4 TTAGGG ismétlődésnél. Ezek jelenléte megakadályozza a telomer telomeráz általi elongációját.[27]
Telomerkvadruplexek
A telomerismétlődésekről számos élőlényben kimutatták in vitro a kvadruplexek létrehozását, később pedig kimutatták ugyanezt in vivo.[28][29] A humán telomerismétlődés (mely minden gerincesnél ugyanaz) a TTAGGG számos ismétlődéséből áll, és az így keletkező kvadruplexek 5-8 nm-es gyöngyszerű szerkezetekben fordulhatnak elő, és NMR-, TEM- és röntgenvizsgálatokkal meghatározták.[30] E kvadruplexek keletkezése csökkenti a telomeráz aktivitását, mely a telomerek hosszáért felel, és a rákok 85%-áért felel. Ez egy aktív kutatási célpont gyógyszerek, például a telomesztatin számára.
Nem telomer kvadruplexek
A kvadruplexek nemcsak telomerekben vannak jelen. Humán, csimpánz-, egér- és patkánygenomok elemzése számos pG4 szekvenciát mutatott a telomereken kívül. Sok ilyen kvadruplex volt promoterekben, és ezek a fajok közt állandók maradtak.[6][7] Sok G-kvadruplex található az E. coliban és több száz más mikrobiális genomban. Itt is a gerincesekhez hasonlóan a G-kvadruplexek gyakoriak a promoterekben.[6] Ezenkívül több mint egymilliárd éves G-kvadruplex-lokusz is ismert növényekben és algákban az RNS-polimeráz II nagy alegységének génjében.[31] Bár e tanulmányok G-kvadruplex-mediált génszabályzást jeleztek előre, nem valószínű, hogy minden pG4 in vivo is keletkezik. A c-myc protoonkogén egy génaktivitásért felelős nukleáz-hiperszenzitív régióban G-kvadruplexet alkot.[32][33] További, promoterében G-kvadruplexet tartalmazó gének például a tyúk-β-globin génje, a humán ubikvitin-ligáz RFP2, valamint a c-kit, bcl-2, VEGF, a H-ras és az N-ras.[34][35][36]
Kvadruplexkeletkezési szabályon alapuló genomfelméréseket is végeztek 276 000 humán feltételezett kvadruplexszekvencián, de ezek nem mindegyike alakít in vivo kvadruplexet.[37] Hasonló tanulmányok azonosítottak feltételezett G-kvadruplexeket prokariótákban, például E. coliban.[38] Számos lehetséges modell van a kvadruplexek génaktivitás-módosítására. Egy modell látható lejjebb, ahol egy promoterben vagy ahhoz közel G-kvadruplex alakul ki, blokkolva a gén transzkripcióját, inaktívvá téve azt. Egy másik modellben a nem kódoló DNS-ben lévő kvadruplex lehetővé teszi a kódoló DNS nyitott állását és a megfelelő gén kifejeződésének növelését.
Funkció
It has been suggested that quadruplex formation plays a role in immunoglobulin heavy chain switching.[5] As cells have evolved mechanisms for resolving (i.e., unwinding) quadruplexes that form. Quadruplex formation may be potentially damaging for a cell; the helicases WRN and Bloom syndrome protein have a high affinity for resolving DNA G-quadruplexes.[39] The DEAH/RHA helicase, DHX36, has also been identified as a key G-quadruplex resolvase.[40][41] In 2009, a metastasis suppressor protein NM23H2 (also known as NME2) was found to directly interact with G-quadruplex in the promoter of the c-myc gene, and transcriptionally regulate c-myc.[42][43] More recently, NM23H2 was reported to interact with G-quadruplex in the promoter of the human telomerase (hTERT) gene and regulate hTERT expression [44] In 2019, the telomere-binding-factor-2 (TRF2 or TERF2) was shown to bind to thousands of non-telomeric G-quadruplexes in the human genome by TRF2 ChIP-seq.[45] There are many studies that implicate quadruplexes in both positive and negative transcriptional regulation, including epigenetic regulation of genes like hTERT.[44] Function of G-quadruplexes have also been reported in allowing programmed recombination of immunologlobin heavy genes and the pilin antigenic variation system of the pathogenic Neisseria.[46] The roles of quadruplex structure in translation control are not as well explored. The direct visualization of G-quadruplex structures in human cells[47] as well as the co-crystal structure of an RNA helicase bound to a G-quadruplex[48] have provided important confirmations of their relevance to cell biology. The potential positive and negative roles of quadruplexes in telomere replication and function remains controversial. T-loops and G-quadruplexes are described as the two tertiary DNA structures that protect telomere ends and regulate telomere length.[49]
Genomszabályzás G-kvadruplexek kialakulásával
Számos genomszabályzó folyamat kapcsolatban állhat G-kvadruplexek keletkezésével, mivel a purin- vagy pirimidinmentes (AP-helyek) javításában játszhat szerepet.[50] A new technique to map AP sites has been developed known as AP-seq which utilizes a biotin-labeled aldehyde-reactive probe (ARP) to tag certain regions of the genome where AP site damage occurrence has been significant.[51] Another genome-wide mapping sequencing method known as ChIP-sequencing, was utilized to map both; damage in AP sites, and the enzyme responsible for its repair, AP endonuclease 1 (APE1). Both of these genome-wide mapping sequencing methods, ChIP-sequencing and ARP, have indicated that AP site damage occurrence is nonrandom. AP site damage was also more prevalent in certain regions of the genome that contain specific active promoter and enhancer markers, some of which were linked to regions responsible for lung adenocarcinoma and colon cancer.[52] AP site damage was found to be predominant in PQS regions of the genome, where formation of G-quadruplex structures is regulated and promoted by the DNA repair process, base excision repair (BER).[52] Base excision repair processes in cells have been proved to be reduced with aging as its components in the mitochondria begin to decline, which can lead to the formation of many diseases such as Alzheimer's disease (AD).[53] These G-quadruplex structures are said to be formed in the promoter regions of DNA through superhelicity, which favors the unwinding of the double helical structure of DNA and in turn loops the strands to form G-quadruplex structures in guanine rich regions.[54] The BER pathway is signalled when it indicates an oxidative DNA base damage, where structures like, 8-Oxoguanine-DNA glycosylase 1 (OGG1), APE1 and G-quadruplex play a huge role in its repair. These enzymes participate in BER to repair certain DNA lesions such as 7,8-dihydro-8-oxoguanine (8-oxoG), which forms under oxidative stress to guanine bases.[55]
Az endogén oxidációs DNS-károsodás szerepe a G4-kialakulásban
Guanine (G) bases in G-quadruplex have the lowest redox potential causing it to be more susceptible to the formation of 8-oxoguanine (8-oxoG), an endogenous oxidized DNA base damage in the genome. Due to Guanine having a lower electron reduction potential than the other nucleotides bases,[56]8-oxo-2'-deoxyguanosine (8-oxo-dG), is a known major product of DNA oxidation. Its concentration is used as a measurement of oxidative stress within a cell.[57] When DNA undergoes oxidative damage, a possible structural change in guanine, after ionizing radiation, gives rise to an enol form, 8-OH-Gua. This oxidative product is formed through a tautomeric shift from the original damage guanine, 8-oxo-Gua, and represents DNA damage that causes changes in the structure. This form allows for the base excision repair (BER) enzyme OGG1 to bind and remove the oxidative damage with the help of APE1, resulting in an AP site.[55][53] Moreover, an AP site is a location in DNA that has neither a purine or a pyrimidine base due to DNA damage, they are the most prevalent type of endogenous DNA damage in cells. AP sites can be generated spontaneously or after the cleavage of modified bases, like 8-OH-Gua.[51] The generation of an AP site enables the melting of the duplex DNA to unmask the PQS, adopting a [53] G-quadruplex fold. With the use of genome-wide ChIP-sequencing analyses, cell-based assays, and in vitro biochemical analyses, a connection has been made between oxidized DNA base-derived AP sites, and the formation of the G-quadruplex.[52]
A DNS oxidációjának kapcsolata betegségekkel
Furthermore, the concentration of 8-oxo-dG is a known biomarker of oxidative stress within a cell, and excessive amount of oxidative stress has been linked to carcinogenesis and other diseases.[58] When produced, 8-oxo-dG, has the ability to inactivate OGG1, thus preventing the repair of DNA damage caused by the oxidation of guanine.[52] The possible inactivation allows for un-repaired DNA damages to gather in non-replicating cells, like muscle, and can cause aging as well.[57] Moreover, oxidative DNA damage like 8-oxo-dG contributes to carcinogenesis through the modulation of gene expression, or the induction of mutations.[57] On the condition that 8-oxo-dG is repaired by BER, parts of the repair protein is left behind which can lead to epigenetic alterations, or the modulation of gene expression.[59] Upon insertion of 8-oxo-dG into thymidine kinase gene of humans, it was determine that if 8-oxo-dG was left unchecked and not repaired by BER, it can lead to frequent mutations and eventually carcinogenesis.[52][53]
APE1 role in Gene Regulation
AP endonuclease 1 (APE1) is an enzyme responsible for the promotion and the formation of G-quadruplex structures. APE1 is mainly in charge of repairing damage caused to AP sites through the BER pathway. APE1 is considered to be very crucial as AP site damage is known to be the most recurring type of endogenous damage to DNA.[59] The oxidation of certain purine bases, like guanine, forms oxidized nucleotides that impairs DNA function by mismatching nucleotides in the sequences.[57] This is more common in PQS sequences which form oxidized structures, such as 8-oxoguanine. Once the cell is aware of oxidative stress and damage, it recruits OGG1 to the site, whose main function is to initiate the BER pathway.[52] OGG1 does so by cleaving the oxidized base and thus creating an AP site, primarily through the process of negative superhelicity.[54] This AP site then signals cells to engage APE1 binding, which binds to the open duplex region.[58] The binding of APE1 then plays an important role by stabilizing the formation of G-quadruplex structures in that region. This promotes formation of G-quadruplex structures by the folding of the stand.[60] This looping process brings four bases in close proximity that will be held together by Hoogsteen base pairing. After this stage the APE1 gets acetylated by multiple lysine residues on the chromatin, forming acetylated APE1 (AcAPE1).[60] AcAPE1 is very crucial to the BER pathway as it acts as a transcriptional coactivator or corepressor, functioning to load transcription factors (TF) into the site of damage allowing it to regulate the gene expression.[61] AcAPE1 is also very important since it allows APE1 to bind for longer periods of time by delay of its dissociation from the sequence, allowing the repair process to be more efficient.[62] Deacetylation of AcAPE1 is the driving force behind the loading of these TFs, where APE1 dissociates from the G-quadruplex structures.[63] When a study downregulated the presence of APE1 and AcAPE1 in the cell, the formation of G-quadruplex structures was inhibited, which proves the importance of APE1 for the formation of these structures. However, not all G-quadruplex structures require APE1 for formation, in fact some of them formed greater G-quadruplex structures in its absence.[52] Therefore, we can conclude that APE1 has two important roles in genome regulation- Stabilizing the formation of g-quadruplex structures and loading the transcriptional factors onto the AP site
Cancer
Telomeres
G-quadruplex forming sequences are prevalent in eukaryotic cells, especially in telomeres, 5` untranslated strands, and translocation hot spots. G-quadruplexes can inhibit normal cell function, and in healthy cells, are easily and readily unwound by helicase. However, in cancer cells that have mutated helicase these complexes cannot be unwound and leads to potential damage of the cell. This causes replication of damaged and cancerous cells. For therapeutic advances, stabilizing the G-quadruplexes of cancerous cells can inhibit cell growth and replication leading to the cell's death.[64]
Promoter Regions
Along with the association of G-quadruplexes in telomeric regions of DNA, G-quadruplex structures have been identified in various human proto-oncogene promoter regions. The structures most present in the promoter regions of these oncogenes tend to be parallel-stranded G-quadruplex DNA structures.[65] Some of these oncogenes include c-KIT, PDGF-A, c-Myc and VEGF, showing the importance of this secondary structure in cancer growth and development. While the formation of G-quadruplex structure vary to some extent for the different promoter regions of oncogenes, the consistent stabilization of these structures have been found in cancer development.[66] Current therapeutic research actively focuses on targeting this stabilization of G-quadruplex structures to arrest unregulated cell growth and division.
One particular gene region, the c-myc pathway, plays an integral role in the regulation of a protein product, c-Myc. With this product, the c-Myc protein functions in the processes of apoptosis and cell growth or development and as a transcriptional control on human telomerase reverse transcriptase.[67] Interaction of c-Myc promoter G-quadruplex with NM23H2 was shown to regulate c-Myc in cancer cells in 2009 [42]
Regulation of c-myc through Human telomerase reverse transcriptase (hTERT) is also directly regulated through promoter G-quadruplex by interaction with the transcription factor NM23H2 where epigenetic modifications were dependent on NM23H2-G-quadruplex association.[44] Recently, hTERT epigenetic regulation reported to be mediated through interaction of hTERT promoter G-quadruplex with the telomeric factor TRF2.[68]
Another gene pathway deals with the VEGF gene, Vascular Endothelial Growth Factor, which remains involved in the process of angiogenesis or the formation of new blood vessels. The formation of an intramolecular G-quadruplex structure has been shown through studies on the polypurine tract of the promoter region of the VEGF gene. Through recent research on the role of G-quadruplex function in vivo, the stabilization of G-quadruplex structures was shown to regulate VEGF gene transcription, with inhibition of transcription factors in this pathway. The intramolecular G-quadruplex structures are formed mostly through the abundant guanine sequence in the promoter region of this specific pathway.[69] The cyclin-dependent cell cycle checkpoint kinase inhibitor-1 CDKN1A (also known as p21) gene harbours promoter G-quadruplex. Interaction of this G-quadruplex with TRF2 (also known as TERF2) resulted in epigenetic regulation of p21, which was tested using the G-quadruplex-binding ligand 360A.[70]
Hypoxia inducible factor 1ɑ, HIF-1ɑ, remains involved in cancer signaling through its binding to Hypoxia Response Element, HRE, in the presence of hypoxia to begin the process of angiogenesis. Through recent research into this specific gene pathway, the polypurine and polypyrimidine region allows for the transcription of this specific gene and the formation of an intramolecular G-quadruplex structure. However, more research is necessary to determine whether the formation of G-quadruplex regulates the expression of this gene in a positive or negative manner.[71]
The c-kit oncogene deals with a pathway that encodes an RTK, which was shown to have elevated expression levels in certain types of cancer. The rich guanine sequence of this promoter region has shown the ability to form a variety of quadruplexes. Current research on this pathway is focusing on discovering the biological function of this specific quadruplex formation on the c-kit pathway, while this quadruplex sequence has been noticed in various species.[36]
The RET oncogene functions in the transcription of kinase which has been abundant in certain types of cancer. The guanine rich sequence in the promoter region for this pathway exudes a necessity for baseline transcription of this receptor tyrosine kinase. In certain types of cancers, the RET protein has shown increased expression levels. The research on this pathway suggested the formation of a G-quadruplex in the promoter region and an applicable target for therapeutic treatments.[72]
Another oncogene pathway involving PDGF-A, platelet-derived growth factor, involves the process of wound healing and function as mitogenic growth factors for cells. High levels of expression of PDGF have been associated with increased cell growth and cancer. The presence of a guanine-rich sequence in the promoter region of PDGF-A has exhibited the ability to form intramolecular parallel G-quadruplex structures and remains suggested to play a role in transcriptional regulation of PDGF-A. However, research has also identified the presence of G-quadruplex structures within this region due to the interaction of TMPyP4 with this promoter sequence.[73]
Therapeutics
Telomeres are generally made up of G-quadruplexes and remain important targets for therapeutic research and discoveries. These complexes have a high affinity for porphyrin rings which makes them effective anticancer agents. However, TMPyP4 has been limited for used due to its non-selectivity toward cancer cell telomeres and normal double stranded DNA (dsDNA). To address this issue analog of TMPyP4, it was synthesized known as 5Me which targets only G quadruplex DNA which inhibits cancer growth more effectively than TMPyP4.[74]
Ligand design and development remains an important field of research into therapeutic reagents due to the abundance of G-quadruplexes and their multiple conformational differences. One type of ligand involving a Quindoline derivative, SYUIQ-05, utilizes the stabilization of G-quadruplexes in promoter regions to inhibit the production of both the c-Myc protein product and the human telomerase reverse transcriptase (hTERT). This main pathway of targeting this region results in the lack of telomerase elongation, leading to arrested cell development. Further research remains necessary for the discovery of a single gene target to minimize unwanted reactivity with more efficient antitumor activity.[67]
Kvadruplexkötő ligandumok
One way of inducing or stabilizing G-quadruplex formation is to introduce a molecule which can bind to the G-quadruplex structure. A number of ligands, which can be both small molecules and proteins, can bind to the G-quadruplex. These ligands can be naturally occurring or synthetic. This has become an increasingly large field of research in genetics, biochemistry, and pharmacology.
A kationos porfinok és a telomesztatin be tudnak ágyazódni G-kvadruplexek közé.
The binding of ligands to G-quadruplexes is vital for anti-cancer pursuits because G-quadruplexes are found typically at translocation hot spots. MM41, a ligand that binds selectively for a quadruplex on the BCL-2 promoter, is shaped with a central core and 4 side chains branching sterically out. The shape of the ligand is vital because it closely matches the quadruplex which has stacked quartets and the loops of nucleic acids holding it together. When bound, MM41's central chromophore is situated on top of the 3’ terminal G-quartet and the side chains of the ligand associate to the loops of the quadruplex. The quartet and the chromophore are bound with a π-π bond while the side chains and loops are not bound but are in close proximity. What makes this binding strong is the fluidity in the position of the loops to better associate with the ligand side chains.[75]
TMPyP4, a cationic porphyrin, is a more well known G4 binding ligand that helps to repress c-Myc. The way in which TMPyP4 binds to G4's is similar to MM41, with the ring stacking onto the external G-quartet and side chains associating to the loops of G4's.[76]
When designing ligands to be bound to G-quadruplexes, the ligands have a higher affinity for parallel folded G-quadruplexes. It has been found that ligands with smaller side chains bind better to the quadruplex because smaller ligands have more concentrated electron density. Furthermore, the hydrogen bonds of ligands with smaller side chains are shorter and therefore stronger. Ligands with mobile side chains, ones that are able to rotate around its center chromophore, associate more strongly to G-quadruplexes because conformation of the G4 loops and the ligand side chains can align.[77]
Kvadruplex-előrejelzési módszerek
A kvadruplexképzésre képes szekvenciák azonosítása fontos a szerepük megértésében. Általában egyszerű mintaegyezést használnak erre a szálon belüli kvadruplex képzésére alkalmas szekvenciákra: d(G3+N1-7G3+N1-7G3+N1-7G3+), ahol N tetszőleges nukleobázis (beleértve a guanint).[78] This rule has been widely used in on-line algorithms. Bár e szabály hatékonyan azonosít G-kvadruplex-képző helyeket, azonosíthat ezenkívül triplex-[79] és i-motívumú C-szál-képzésre alkalmas nem tökéletes homopurintükör-ismétlődések egy részét is.[80] Továbbá e szekvenciák képesek csúszott és visszahajtott szerkezetek azonosítására is, melyek kvadruplex-[4] és triplex-DNS képzésére alkalmasak.[81] Egy tanulmányban,[82] S. S. Smith kimutatta, hogy e motívumok bázispáronkénti megfigyelt száma (frekvenciája) a teljes elérhető genomszekvenciával rendelkező Eumetazoa-fajokban nagyobb. Ez alapján e szekvenciák pozitív szelekció alatt lehetnek, melyet a nem B-szerkezet kialakulását megakadályozó rendszerek fejlődése tett lehetővé.
2016-tól fejlett webalapú eszköztárak érhetők el G-kvadruplex-képző szekvenciák azonosítására, például a G4Hunter nyílt hozzáférésű felhasználóbarát, csúszóablakos változata[83] vagy a gépi tanuláson alapuló G4RNA Screener.[84]
Módszerek G-kvadruplexek tanulmányozására
Számos kísérleti módszert fejlesztettek ki G-kvadruplexek azonosítására. Ezek két nagy csoportba (biofizikai és biokémiai) sorolhatók.[85]
Biokémiai
A G-kvadruplex-képződés hosszabb szekvenciában való vizsgálatára biokémiai módszerek használhatók. A DNS-polimeráz stop-assay-ben a G-kvadruplex DNS-templátban való keletkezése a polimeráz megállását okozó akadály lehet, mely a primer meghosszabbítását megállítja.[86] A dimetil-szulfát (DMS)-piperidin bontóassay azon alaul, hogy a G-kvadruplex képződése megakadályozza a DMS általi guaninmetilációt, vvédelmi mintához vezetve a DNS G-kvadruplexénél a piperidines bontás után.[87]
Biofizikai
A G-kvadruplex alakja meghatározható a körkörös dikroizmus jeleinek vizsgálatával adott hullámhosszokon.[88] A párhuzamos G-kvadruplexek 240 nm-es CD-jele negatív, 262 nm-es jele pozitív, az antipárhuzamosak jele 262 nm-en negatív, 295-ön pozitív. A G-kvadruplex-képződés igazolásához a CD-kísérlet nem G-kvadruplex-stabilizáló (Li+), illetve G-kvadruplex-stabilizáló (K+ vagy G-kvadruplex ligandumokkal) és a távoli UV tartományban (180-230 nm) kell vizsgáni. Ehhez hasonlóan a G-kvadruplexek hőstabilitása az UV-jel 295 nm-en való vizsgálatával azonosítható.[89] A G-kvadruplexek olvadásakor az UV-elnyelés 295 nm-en csökken, a G-kvadruplexre jellemző hipokrómos váltást okozva. További megközelítés a G-kvadruplex észlelésére például a nanopórus-alapú módszer. Először kiderült, hogy a biológiai nanopórusok képesek a G-kvadruplexek észlelésére méretalapú kizárás alapján és a G-kvadruplexek és a fehérjék nanoüregei közti specifikus interakciókkal.[90] Egy újabb megközeítésbem a szilárdtest-nanopórusok és a DNS-nanotechnológia használatosak a G-kvadruplexek észleléséhez dsDNS-ben és a G-kvadruplex-keletkezés észlelésére.[91]
Szerepe neurológiai betegségekben
G-quadruplexes have been implicated in neurological disorders through two main mechanisms. The first is through expansions of G-repeats within genes that lead to the formation of G-quadruplex structures that directly cause disease, as is the case with the C9orf72 gene and amyotrophic lateral sclerosis (ALS) or frontotemporal dementia (FTD). The second mechanism is through mutations that affect the expression of G-quadruplex binding proteins, as seen in the fragile X mental retardation gene 1 (FMR1) gene and Fragile X Syndrome.[92]
The C9orf72 gene codes for the protein C9orf72 which is found throughout the brain in neuronal cytoplasm and at presynaptic terminals.[93] Mutations of the C9orf72 gene have been linked to the development of FTD and ALS.[94] These two diseases have a causal relationship to GGGGCC (G4C2) repeats within the 1st intron of C9orf72 gene. Normal individuals typically have around 2 to 8 G4C2 repeats, but individuals with FTD or ALS have from 500 to several thousand G4C2 repeats.[95][96] The transcribed RNA of these repeats have been shown to form stable G-quadruplexes, with evidence showing that the G4C2 repeats in DNA have the ability to form mixed parallel-antiparallel G-quadruplex structures as well.[97][98] These RNA transcripts containing G4C2 repeats were shown to bind and separate a wide variety of proteins, including nucleolin. Nucleolin is involved in the synthesis and maturation of ribosomes within the nucleus, and separation of nucleolin by the mutated RNA transcripts impairs nucleolar function and ribosomal RNA synthesis.[99]
Fragile X mental retardation protein (FMRP) is a widely expressed protein coded by the FMR1 gene that binds to G-quadruplex secondary structures in neurons and is involved in synaptic plasticity.[100] FMRP acts as a negative regulator of translation, and its binding stabilizes G-quadruplex structures in mRNA transcripts, inhibiting ribosome elongation of mRNA in the neuron's dendrite and controlling the timing of the transcript's expression.[101][102] Mutations of this gene can cause the development of Fragile X Syndrome, autism, and other neurological disorders.[103] Specifically, Fragile X Syndrome is caused by an increase from 50 to over 200 CGG repeats within exon 13 of the FMR1 gene. This repeat expansion promotes DNA methylation and other epigenetic heterochromatin modifications of FMR1 that prevent the transcription of the gene, leading to pathological low levels of FMRP.[104][105]
Terápiás megközelítések
Antisense-mediated interventions and small-molecule ligands are common strategies used to target neurological diseases linked to G-quadruplex expansion repeats. Therefore, these techniques are especially advantageous for targeting neurological diseases that have a gain-of-function mechanism, which is when the altered gene product has a new function or new expression of a gene; this has been detected in the C9orf72 (chromosome 9 open reading frame 72).[106]
Az antiszensz-terápia során szintetizált nukleinsavszálak kapcsolódnak közvetlenül és specifikusan egy gén mRNS-éhez, inaktiválva azt. Az antiszensz oligonukleotidok (ASO) gyakoriak a C9orf72 RNS célzására a GGGGCC ismétlődő szakaszban, csökkentve a C9orf72 sejtmodelljeiben a toxicitást.[107][108][109] ASOs have previously been used to restore normal phenotypes in other neurological diseases that have gain-of-function mechanisms, the only difference is that it was used in the absence of G-quadruplex expansion repeat regions.[110][111][112][113]
The G-quadruplex decoy strategy is another promising approach for targeting cancer cells by exploiting the unique structural features of the G-quadruplex. The strategy involves designing synthetic oligonucleotides that mimic the G-quadruplex structure and compete with the endogenous G-quadruplexes for binding to transcription factors. These decoys are typically composed of a G-rich sequence that can form a stable G-quadruplex structure and a short linker region that can be modified to optimize their properties.[114] When introduced to cancer cells the decoy can intercept associated transcription factors and bind them leading to the regulation of gene expression. Decoys have been successfully demonstrated to inhibit oncogenic KRAS in SCID mice leading to reduced tumour growth and increased median survival time.[115]
Another commonly used technique is the utilization of small-molecule ligands. These can be used to target G-quadruplex regions that cause neurological disorders. Approximately 1,000 various G-quadruplex ligands exist in which they are able to interact via their aromatic rings; this allows the small-molecule ligands to stack on the planar terminal tetrads within the G-quadruplex regions. A disadvantage of using small-molecule ligands as a therapeutic technique is that specificity is difficult to manage due to the variability of G-quadruplexes in their primary sequences, orientation, thermodynamic stability, and nucleic acid strand stoichiometry. 2020. márciusáig nem ismert egy kis ligandum, mely egy adott G-kvadruplexre lenne specifikus.[116][117] However, a cationic porphyrin known as TMPyP4 is able to bind to the C9orf72 GGGGCC repeat region, which causes the G-quadruplex repeat region to unfold and lose its interactions with proteins causing it to lose its functionality.[118] Small-molecule ligands, composed primarily of lead, can target GGGGCC repeat regions as well and ultimately decreased both repeat-associated non-ATG translation and RNA foci in neuron cells derived from patients with Amyotrophic lateral sclerosis (ALS). This provides evidence that small-molecule ligands are an effective and efficient process to target GGGGCC regions, and that specificity for small-molecule ligand binding is a feasible goal for the scientific community.
A fémkomplexek számos G4-DNS-kötésre alkalmassá és lehetséges gyógyszerré tevő jellemzővel rendelkeznek. Míg a fém főleg szerkezeti szereppel rendelkezik a legtöbb G4-kötőben, vannak esetek, mikor közvetlenül a G4-gyel lépnek kölcsönhatásba elektrosztatikusan vagy a nukleobázisokkal való közvetlen koordinációval.[119]
Jegyzetek
- ↑ J. A. Capra, K. Paeschke, M. Singh, V. A. Zakian (2010. július 1.). „G-quadruplex DNA sequences are evolutionarily conserved and associated with distinct genomic features in Saccharomyces cerevisiae”. PLOS Computational Biology 6 (7), e1000861. o. DOI:10.1371/journal.pcbi.1000861. PMID 20676380.
- ↑ (2017. március 1.) „A G-quadruplex DNA-affinity Approach for Purification of Enzymaticacvly Active G4 Resolvase1”. Journal of Visualized Experiments 121 (121). DOI:10.3791/55496. PMID 28362374.
- ↑ a b E. Largy, J. Mergny, V. Gabelica.szerk.: S. Astrid, S. Helmut, K. O. Roland: Chapter 7. Role of Alkali Metal Ions in G-Quadruplex Nucleic Acid Structure and Stability, The Alkali Metal Ions: Their Role in Life, Metal Ions in Life Sciences. Springer, 203–258. o.. DOI: 10.1007/978-3-319-21756-7_7 (2016)
- ↑ a b W. I. Sundquist, A. Klug (1989. december 1.). „Telomeric DNA dimerizes by formation of guanine tetrads between hairpin loops”. Nature 342 (6251), 825–829. o. DOI:10.1038/342825a0. PMID 2601741.
- ↑ a b c (1988. július 1.) „Formation of parallel four-stranded complexes by guanine-rich motifs in DNA and its implications for meiosis”. Nature 334 (6180), 364–366. o. DOI:10.1038/334364a0. PMID 3393228.
- ↑ a b c d (2006. május 3.) „Genome-wide Prediction of G4 DNA as Regulatory Motifs: Role in Escherichia Coli Global Regulation”. Genome Research 16 (5), 644‐655. o. DOI:10.1101/gr.4508806. PMID 16651665.
- ↑ a b c d S. Borman (2007. május 28.). „Ascent of quadruplexes nucleic acid structures become promising drug targets”. Chemical and Engineering News 85 (22), 12–17. o. DOI:10.1021/cen-v085n009.p012a.
- ↑ (2008) „Genome-wide Computational and Expression Analyses Reveal G-quadruplex DNA Motifs as Conserved Cis-Regulatory Elements in Human and Related Species”. Journal of Medicinal Chemistry 51 (18), 5641‐5649. o. DOI:10.1021/jm800448a. PMID 18767830.
- ↑ H. Han, L. H. Hurley (2000. április 1.). „G-quadruplex DNA: a potential target for anti-cancer drug design”. Trends in Pharmacological Sciences 21 (4), 136–142. o. DOI:10.1016/s0165-6147(00)01457-7. PMID 10740289.
- ↑ M. L. Bochman, K. Paeschke, V. A. Zakian (2012. november 1.). „DNA secondary structures: stability and function of G-quadruplex structures”. Nature Reviews. Genetics 13 (11), 770–80. o. DOI:10.1038/nrg3296. PMID 23032257.
- ↑ (2008. május 3.) „QuadBase: Genome-Wide Database of G4 DNA--occurrence and Conservation in Human, Chimpanzee, Mouse and Rat Promoters and 146 Microbes”. Nucleic Acids Research 36 (Database), D381‐D385. o. DOI:10.1093/nar/gkm781. PMID 17962308.
- ↑ P. Dhapola, S. Chowdhury (2016. július 1.). „QuadBase2: Web Server for Multiplexed Guanine Quadruplex Mining and Visualization”. Nucleic Acids Research 44 (W1), W277‐W283. o. DOI:10.1093/nar/gkw425. PMID 27185890.
- ↑ D. Rhodes, H. J. Lipps (2015. október 1.). „G-quadruplexes and their regulatory roles in biology”. Nucleic Acids Research 43 (18), 8627–37. o. DOI:10.1093/nar/gkv862. PMID 26350216.
- ↑ S. Borman (2009. november 1.). „Promoter quadruplexes folded DNA structures in gene-activation sites may be useful cancer drug targets”. Chemical and Engineering News 87 (44), 28–30. o. DOI:10.1021/cen-v087n044.p028.
- ↑ (1962. december 1.) „Helix formation by guanylic acid”. Proceedings of the National Academy of Sciences of the United States of America 48 (12), 2013–2018. o. DOI:10.1073/pnas.48.12.2013. PMID 13947099.
- ↑ (1987. december 1.) „Telomeric DNA oligonucleotides form novel intramolecular structures containing guanine-guanine base pairs”. Cell 51 (6), 899–908. o. DOI:10.1016/0092-8674(87)90577-0. PMID 3690664.
- ↑ (2010. október 10.) „Small-molecule-mediated G-quadruplex isolation from human cells”. Nature Chemistry 2 (12), 1095–1098. o. DOI:10.1038/nchem.842. PMID 21107376.
- ↑ (2012. február 5.) „Small-molecule–induced DNA damage identifies alternative DNA structures in human genes”. Nature Chemical Biology 8 (3), 301–310. o. DOI:10.1038/nchembio.780. PMID 22306580.
- ↑ T. Simonsson (2001. április 1.). „G-quadruplex DNA structures--variations on a theme”. Biological Chemistry 382 (4), 621–8. o. DOI:10.1515/BC.2001.073. PMID 11405224.
- ↑ (2006) „Quadruplex DNA: sequence, topology and structure”. Nucleic Acids Research 34 (19), 5402–15. o. DOI:10.1093/nar/gkl655. PMID 17012276.
- ↑ (2012. május 1.) „Computational detection and analysis of sequences with duplex-derived interstrand G-quadruplex forming potential”. Methods 57 (1), 3–10. o. DOI:10.1016/j.ymeth.2012.05.002. PMID 22652626.
- ↑ (2016) „G-Quadruplexes Involving Both Strands of Genomic DNA Are Highly Abundant and Colocalize with Functional Sites in the Human Genome”. PLOS ONE 11 (1), e0146174. o. DOI:10.1371/journal.pone.0146174. PMID 26727593.
- ↑ (2014. április 1.) „Existence and consequences of G-quadruplex structures in DNA”. Current Opinion in Genetics & Development 25 (25), 22–29. o. DOI:10.1016/j.gde.2013.10.012. PMID 24584093.
- ↑ (2006. június 1.) „Hydration regulates thermodynamics of G-quadruplex formation under molecular crowding conditions”. Journal of the American Chemical Society 128 (24), 7957–63. o. DOI:10.1021/ja061267m. PMID 16771510.
- ↑ (2010. január 1.) „Molecular crowding creates an essential environment for the formation of stable G-quadruplexes in long double-stranded DNA”. Nucleic Acids Research 38 (1), 327–38. o. DOI:10.1093/nar/gkp898. PMID 19858105.
- ↑ (2016. február 1.) „Real-Time Monitoring of G-Quadruplex Formation during Transcription”. Analytical Chemistry 88 (4), 1984–9. o. DOI:10.1021/acs.analchem.5b04396. PMID 26810457.
- ↑ (2011. augusztus 1.) „G-quadruplex formation at the 3' end of telomere DNA inhibits its extension by telomerase, polymerase and unwinding by helicase”. Nucleic Acids Research 39 (14), 6229–37. o. DOI:10.1093/nar/gkr164. PMID 21441540.
- ↑ (2001. július 1.) „In vitro generated antibodies specific for telomeric guanine-quadruplex DNA react with Stylonychia lemnae macronuclei”. Proceedings of the National Academy of Sciences of the United States of America 98 (15), 8572–7. o. DOI:10.1073/pnas.141229498. PMID 11438689.
- ↑ (2005. október 1.) „Telomere end-binding proteins control the formation of G-quadruplex DNA structures in vivo”. Nature Structural & Molecular Biology 12 (10), 847–54. o. DOI:10.1038/nsmb982. PMID 16142245.
- ↑ (2018. június 1.) „Long repeating (TTAGGG)n single-stranded DNA self-condenses into compact beaded filaments stabilized by G-quadruplex formation”. The Journal of Biological Chemistry 293 (24), 9473–9485. o. DOI:10.1074/jbc.RA118.002158. PMID 29674319.
- ↑ (2021. július 1.) „G-Quadruplex in Gene Encoding Large Subunit of Plant RNA Polymerase II: A Billion-Year-Old Story”. International Journal of Molecular Sciences 22 (14), 7381. o. DOI:10.3390/ijms22147381. PMID 34299001.
- ↑ (1998. március 1.) „DNA tetraplex formation in the control region of c-myc”. Nucleic Acids Research 26 (5), 1167–72. o. DOI:10.1093/nar/26.5.1167. PMID 9469822.
- ↑ (2002. szeptember 1.) „Direct evidence for a G-quadruplex in a promoter region and its targeting with a small molecule to repress c-MYC transcription”. Proceedings of the National Academy of Sciences of the United States of America 99 (18), 11593–8. o. DOI:10.1073/pnas.182256799. PMID 12195017.
- ↑ (2006. december 14.) „G-quadruplexes in promoters throughout the human genome”. Nucleic Acids Research 35 (2), 406–13. o. DOI:10.1093/nar/gkl1057. PMID 17169996.
- ↑ (2006. február 1.) „An intramolecular G-quadruplex structure with mixed parallel/antiparallel G-strands formed in the human BCL-2 promoter region in solution”. Journal of the American Chemical Society 128 (4), 1096–8. o. DOI:10.1021/ja055636a. PMID 16433524.
- ↑ a b (2006. június 1.) „A conserved quadruplex motif located in a transcription activation site of the human c-kit oncogene”. Biochemistry 45 (25), 7854–60. o. DOI:10.1021/bi0601510. PMID 16784237.
- ↑ (2005) „Prevalence of quadruplexes in the human genome”. Nucleic Acids Research 33 (9), 2908–16. o. DOI:10.1093/nar/gki609. PMID 15914667.
- ↑ (2006. május 1.) „Genome-wide prediction of G4 DNA as regulatory motifs: role in Escherichia coli global regulation”. Genome Research 16 (5), 644–55. o. DOI:10.1101/gr.4508806. PMID 16651665.
- ↑ (2012) „The Werner syndrome protein is distinguished from the Bloom syndrome protein by its capacity to tightly bind diverse DNA structures”. PLOS ONE 7 (1), e30189. o. DOI:10.1371/journal.pone.0030189. PMID 22272300.
- ↑ (2005. november 1.) „The DEXH protein product of the DHX36 gene is the major source of tetramolecular quadruplex G4-DNA resolving activity in HeLa cell lysates”. The Journal of Biological Chemistry 280 (46), 38117–20. o. DOI:10.1074/jbc.C500348200. PMID 16150737.
- ↑ (2017. augusztus 15.) „Structural Basis of DEAH/RHA Helicase Activity”. Crystals 7 (8), 253. o. DOI:10.3390/cryst7080253.
- ↑ a b (2009. január 1.) „Metastases Suppressor NM23-H2 Interaction With G-quadruplex DNA Within c-MYC Promoter Nuclease Hypersensitive Element Induces c-MYC Expression”. Nucleic Acids Research 37 (1), 172‐183. o. DOI:10.1093/nar/gkn919. PMID 19033359.
- ↑ (2009. november 1.) „Promoter Quadruplexes Folded DNA structures in gene-activation sites may be useful cancer drug targets”. Chemical and Engineering News 87 (44), 28–30. o. DOI:10.1021/cen-v087n044.p028.
- ↑ a b c (2017. július 1.) „Epigenetic Suppression of Human Telomerase ( hTERT) Is Mediated by the Metastasis Suppressor NME2 in a G-quadruplex-dependent Fashion”. The Journal of Biological Chemistry 292 (37), 15205‐15215. o. DOI:10.1074/jbc.M117.792077. PMID 28717007.
- ↑ (2019. november 1.) „Telomere Repeat-Binding Factor 2 Binds Extensively to Extra-Telomeric G-quadruplexes and Regulates the Epigenetic Status of Several Gene Promoters”. The Journal of Biological Chemistry 294 (47), 17709–17722. o. DOI:10.1074/jbc.RA119.008687. PMID 31575660.
- ↑ (2013. április 1.) „The G4 genome”. PLOS Genetics 9 (4), e1003468. o. DOI:10.1371/journal.pgen.1003468. PMID 23637633.
- ↑ (2013. március 1.) „Quantitative visualization of DNA G-quadruplex structures in human cells”. Nature Chemistry 5 (3), 182–6. o. DOI:10.1038/nchem.1548. PMID 23422559.
- ↑ (2018. június 1.) „Structural basis of G-quadruplex unfolding by the DEAH/RHA helicase DHX36” (angol nyelven). Nature 558 (7710), 465–469. o. DOI:10.1038/s41586-018-0209-9. PMID 29899445.
- ↑ (2016) „Structure and function of the telomeric CST complex”. Computational and Structural Biotechnology Journal 14, 161–7. o. DOI:10.1016/j.csbj.2016.04.002. PMID 27239262.
- ↑ (2016. október 1.) „G-quadruplex structures mark human regulatory chromatin”. Nature Genetics 48 (10), 1267–1272. o. DOI:10.1038/ng.3662. PMID 27618450.
- ↑ a b AP-Seq: A Method to Measure Apurinic Sites and Small Base Adducts Genome-Wide, The Nucleus, Methods in Molecular Biology, 95–108. o.. DOI: 10.1007/978-1-0716-0763-3_8 (2020. május 3.). ISBN 978-1-0716-0762-6
- ↑ a b c d e f g (2020. május 1.) „Endogenous oxidized DNA bases and APE1 regulate the formation of G-quadruplex structures in the genome”. Proceedings of the National Academy of Sciences of the United States of America 117 (21), 11409–11420. o. DOI:10.1073/pnas.1912355117. PMID 32404420.
- ↑ a b c d (2014. június 1.) „Base excision DNA repair levels in mitochondrial lysates of Alzheimer's disease”. Neurobiology of Aging 35 (6), 1293–1300. o. DOI:10.1016/j.neurobiolaging.2014.01.004. PMID 24485507.
- ↑ a b (2009. május 1.) „The importance of negative superhelicity in inducing the formation of G-quadruplex and i-motif structures in the c-Myc promoter: implications for drug targeting and control of gene expression”. Journal of Medicinal Chemistry 52 (9), 2863–2874. o. DOI:10.1021/jm900055s. PMID 19385599.
- ↑ a b (2001. január 1.) „Stimulation of human 8-oxoguanine-DNA glycosylase by AP-endonuclease: potential coordination of the initial steps in base excision repair”. Nucleic Acids Research 29 (2), 430–438. o. DOI:10.1093/nar/29.2.430. PMID 11139613.
- ↑ (1998. május 1.) „Oxidative Nucleobase Modifications Leading to Strand Scission”. Chemical Reviews 98 (3), 1109–1152. o. DOI:10.1021/cr960421s. PMID 11848927.
- ↑ a b c d (2020. január 7.) „The genomics of oxidative DNA damage, repair, and resulting mutagenesis”. Computational and Structural Biotechnology Journal 18, 207–219. o. DOI:10.1016/j.csbj.2019.12.013. PMID 31993111.
- ↑ a b (2017. október 1.) „8-Oxo-7,8-dihydro-2'-deoxyguanosine and abasic site tandem lesions are oxidation prone yielding hydantoin products that strongly destabilize duplex DNA”. Organic & Biomolecular Chemistry 15 (39), 8341–8353. o. DOI:10.1039/C7OB02096A. PMID 28936535.
- ↑ a b (2019. szeptember 1.) „Nucleotide excision repair of abasic DNA lesions”. Nucleic Acids Research 47 (16), 8537–8547. o. DOI:10.1093/nar/gkz558. PMID 31226203.
- ↑ a b (2017. március 1.) „Human Apurinic/Apyrimidinic Endonuclease (APE1) Is Acetylated at DNA Damage Sites in Chromatin, and Acetylation Modulates Its DNA Repair Activity”. Molecular and Cellular Biology 37 (6). DOI:10.1128/mcb.00401-16. PMID 27994014.
- ↑ (2008. december 1.) „Regulatory role of human AP-endonuclease (APE1/Ref-1) in YB-1-mediated activation of the multidrug resistance gene MDR1”. Molecular and Cellular Biology 28 (23), 7066–7080. o. DOI:10.1128/mcb.00244-08. PMID 18809583.
- ↑ (2003. december 1.) „Role of acetylated human AP-endonuclease (APE1/Ref-1) in regulation of the parathyroid hormone gene”. The EMBO Journal 22 (23), 6299–6309. o. DOI:10.1093/emboj/cdg595. PMID 14633989.
- ↑ (2010. január 1.) „SIRT1 deacetylates APE1 and regulates cellular base excision repair”. Nucleic Acids Research 38 (3), 832–845. o. DOI:10.1093/nar/gkp1039. PMID 19934257.
- ↑ (2016. július 1.) „Quadruplex Nucleic Acids as Novel Therapeutic Targets” (angol nyelven). Journal of Medicinal Chemistry 59 (13), 5987–6011. o. DOI:10.1021/acs.jmedchem.5b01835. PMID 26840940.
- ↑ Sequence, stability, and structure of G-quadruplexes and their interactions with drugs, 17.5.1–17.5.17. o.. DOI: 10.1002/0471142700.nc1705s50 (2012. szeptember 1.). ISBN 978-0471142706
- ↑ (2010. szeptember 1.) „Making sense of G-quadruplex and i-motif functions in oncogene promoters”. The FEBS Journal 277 (17), 3459–69. o. DOI:10.1111/j.1742-4658.2010.07759.x. PMID 20670278.
- ↑ a b (2011. augusztus 1.) „Inhibition of cell proliferation by quindoline derivative (SYUIQ-05) through its preferential interaction with c-myc promoter G-quadruplex”. Journal of Medicinal Chemistry 54 (16), 5671–9. o. DOI:10.1021/jm200062u. PMID 21774525.
- ↑ (2020. január 1.) „Human Telomerase Expression is under Direct Transcriptional Control of the Telomere-binding-factor TRF2”. bioRxiv. DOI:10.1101/2020.01.15.907626.
- ↑ (2005. október 12.) „Facilitation of a structural transition in the polypurine/polypyrimidine tract within the proximal promoter region of the human VEGF gene by the presence of potassium and G-quadruplex-interactive agents”. Nucleic Acids Research 33 (18), 6070–80. o. DOI:10.1093/nar/gki917. PMID 16239639.
- ↑ (2017. szeptember 1.) „Transcription Regulation of CDKN1A (p21/CIP1/WAF1) by TRF2 Is Epigenetically Controlled Through the REST Repressor Complex”. Scientific Reports 7 (1), 11541. o. DOI:10.1038/s41598-017-11177-1. PMID 28912501.
- ↑ (2005. december 1.) „Evidence for the presence of a guanine quadruplex forming region within a polypurine tract of the hypoxia inducible factor 1alpha promoter”. Biochemistry 44 (49), 16341–50. o. DOI:10.1021/bi051618u. PMID 16331995.
- ↑ (2007. augusztus 1.) „Formation of pseudosymmetrical G-quadruplex and i-motif structures in the proximal promoter region of the RET oncogene”. Journal of the American Chemical Society 129 (33), 10220–8. o. DOI:10.1021/ja072185g. PMID 17672459.
- ↑ (2007. november 26.) „Characterization of the G-quadruplexes in the duplex nuclease hypersensitive element of the PDGF-A promoter and modulation of PDGF-A promoter activity by TMPyP4”. Nucleic Acids Research 35 (22), 7698–713. o. DOI:10.1093/nar/gkm538. PMID 17984069.
- ↑ (2018. november 1.) „Novel Amphiphilic G-Quadruplex Binding Synthetic Derivative of TMPyP4 and Its Effect on Cancer Cell Proliferation and Apoptosis Induction”. Biochemistry 57 (46), 6514–6527. o. DOI:10.1021/acs.biochem.8b00843. PMID 30369235.
- ↑ (2015. június 1.) „A G-quadruplex-binding compound showing anti-tumour activity in an in vivo model for pancreatic cancer”. Scientific Reports 5, 11385. o. DOI:10.1038/srep11385. PMID 26077929.
- ↑ (2002. szeptember 1.) „Direct evidence for a G-quadruplex in a promoter region and its targeting with a small molecule to repress c-MYC transcription”. Proceedings of the National Academy of Sciences of the United States of America 99 (18), 11593–8. o. DOI:10.1073/pnas.182256799. PMID 12195017.
- ↑ (2012. február 1.) „Structural basis for telomeric G-quadruplex targeting by naphthalene diimide ligands” (angol nyelven). Journal of the American Chemical Society 134 (5), 2723–31. o. DOI:10.1021/ja2102423. PMID 22280460.
- ↑ (2005) „Highly prevalent putative quadruplex sequence motifs in human DNA”. Nucleic Acids Research 33 (9), 2901–7. o. DOI:10.1093/nar/gki553. PMID 15914666.
- ↑ (1995) „Triplex DNA structures”. Annual Review of Biochemistry 64 (9), 65–95. o. DOI:10.1146/annurev.bi.64.070195.000433. PMID 7574496.
- ↑ (2008. augusztus 1.) „Intramolecularly folded G-quadruplex and i-motif structures in the proximal promoter of the vascular endothelial growth factor gene”. Nucleic Acids Research 36 (14), 4598–608. o. DOI:10.1093/nar/gkn380. PMID 18614607.
- ↑ (1987) „DNA H form requires a homopurine-homopyrimidine mirror repeat”. Nature 330 (6147), 495–7. o. DOI:10.1038/330495a0. PMID 2825028.
- ↑ S. S. Smith (2010). „Evolutionary expansion of structurally complex DNA sequences”. Cancer Genomics & Proteomics 7 (4), 207–15. o. PMID 20656986.
- ↑ (2019. szeptember 15.) „G4Hunter web application: a web server for G-quadruplex prediction” (angol nyelven). Bioinformatics 35 (18), 3493–3495. o. DOI:10.1093/bioinformatics/btz087. ISSN 1367-4803. PMID 30721922.
- ↑ (2018) „G4RNA screener web server: User focused interface for RNA G-quadruplex prediction” (angol nyelven). Biochimie 151, 115–118. o. DOI:10.1016/j.biochi.2018.06.002. PMID 29885355.
- ↑ C. K. Kwok, C. J. Merrick (2017. október 1.). „G-Quadruplexes: Prediction, Characterization, and Biological Application”. Trends in Biotechnology 35 (10), 997–1013. o. DOI:10.1016/j.tibtech.2017.06.012. PMID 28755976.
- ↑ H. Han, L. H. Hurley, M. Salazar (1999. január 1.). „A DNA polymerase stop assay for G-quadruplex-interactive compounds”. Nucleic Acids Research 27 (2), 537–542. o. DOI:10.1093/nar/27.2.537. PMID 9862977.
- ↑ Biochemical Techniques for the Characterization of G-Quadruplex Structures: EMSA, DMS Footprinting, and DNA Polymerase Stop Assay, G-Quadruplex DNA, Methods in Molecular Biology. Humana Press, 65–79. o.. DOI: 10.1007/978-1-59745-363-9_5 (2009. október 23.). ISBN 9781588299505
- ↑ S- Paramasovan, I. Rujan, P. H. Bolton (2007. december 1.). „Circular dichroism of quadruplex DNAs: applications to structure, cation effects and ligand binding”. Methods 43 (4), 324–331. o. DOI:10.1016/j.ymeth.2007.02.009. PMID 17967702.
- ↑ (1998. szeptember 1.) „Following G-quartet formation by UV-spectroscopy”. FEBS Letters 435 (1), 74–78. o. DOI:10.1016/s0014-5793(98)01043-6. PMID 9755862.
- ↑ (2014. október 1.) „Single-molecule investigation of G-quadruplex folds of the human telomere sequence in a protein nanocavity”. Proceedings of the National Academy of Sciences of the United States of America 111 (40), 14325–14331. o. DOI:10.1073/pnas.1415944111. PMID 25225404.
- ↑ (2019. november 1.) „Monitoring G-Quadruplex Formation with DNA Carriers and Solid-State Nanopores”. Nano Letters 19 (11), 7996–8001. o. DOI:10.1021/acs.nanolett.9b03184. PMID 31577148.
- ↑ (2015. június 1.) „G-quadruplexes: Emerging roles in neurodegenerative diseases and the non-coding transcriptome”. FEBS Letters 589 (14), 1653–68. o. DOI:10.1016/j.febslet.2015.05.003. PMID 25979174.
- ↑ C9orf72 chromosome 9 open reading frame 72 [Homo sapiens] - Gene]. National Center for Biotechnology Information. U.S. National Library of Medicine
- ↑ (2002. június 1.) „The prevalence of frontotemporal dementia”. Neurology 58 (11), 1615–21. o. DOI:10.1212/WNL.58.11.1615. PMID 12058088.
- ↑ (2012. december 1.) „Length of normal alleles of C9ORF72 GGGGCC repeat do not influence disease phenotype”. Neurobiology of Aging 33 (12), 2950.e5–7. o. DOI:10.1016/j.neurobiolaging.2012.07.005. PMID 22840558.
- ↑ (2013. március 1.) „Large C9orf72 hexanucleotide repeat expansions are seen in multiple neurodegenerative syndromes and are more frequent than expected in the UK population”. American Journal of Human Genetics 92 (3), 345–53. o. DOI:10.1016/j.ajhg.2013.01.011. PMID 23434116.
- ↑ (2012. december 1.) „C9orf72 hexanucleotide repeat associated with amyotrophic lateral sclerosis and frontotemporal dementia forms RNA G-quadruplexes”. Scientific Reports 2, 1016. o. DOI:10.1038/srep01016. PMID 23264878.
- ↑ (2013. április 1.) „The disease-associated r(GGGGCC)n repeat from the C9orf72 gene forms tract length-dependent uni- and multimolecular RNA G-quadruplex structures”. The Journal of Biological Chemistry 288 (14), 9860–6. o. DOI:10.1074/jbc.C113.452532. PMID 23423380.
- ↑ (2014. március 1.) „C9orf72 nucleotide repeat structures initiate molecular cascades of disease”. Nature 507 (7491), 195–200. o. DOI:10.1038/nature13124. PMID 24598541.
- ↑ (2001. november 1.) „Fragile X mental retardation protein targets G quartet mRNAs important for neuronal function”. Cell 107 (4), 489–499. o. DOI:10.1016/S0092-8674(01)00566-9. PMID 11719189.
- ↑ (2003. december 1.) „Phosphorylation influences the translation state of FMRP-associated polyribosomes”. Human Molecular Genetics 12 (24), 3295–3305. o. DOI:10.1093/hmg/ddg350. PMID 14570712.
- ↑ (2009. február 1.) „Translational regulation of the human achaete-scute homologue-1 by fragile X mental retardation protein”. The Journal of Biological Chemistry 284 (7), 4255–4266. o. DOI:10.1074/jbc.M807354200. PMID 19097999.
- ↑ Fragile X Mental Retardation. Gene Cards
- ↑ (1991. augusztus 1.) „Absence of expression of the FMR-1 gene in fragile X syndrome”. Cell 66 (4), 817–822. o. DOI:10.1016/0092-8674(91)90125-I. PMID 1878973.
- ↑ (1992. szeptember 1.) „DNA methylation represses FMR-1 transcription in fragile X syndrome”. Human Molecular Genetics 1 (6), 397–400. o. DOI:10.1093/hmg/1.6.397. PMID 1301913.
- ↑ (2014. október 1.) „C9orf72 amyotrophic lateral sclerosis and frontotemporal dementia: gain or loss of function?”. Current Opinion in Neurology 27 (5), 515–23. o. DOI:10.1097/WCO.0000000000000130. PMID 25188012.
- ↑ (2013. október 1.) „RNA toxicity from the ALS/FTD C9ORF72 expansion is mitigated by antisense intervention”. Neuron 80 (2), 415–28. o. DOI:10.1016/j.neuron.2013.10.015. PMID 24139042.
- ↑ (2013. november 1.) „Targeted degradation of sense and antisense C9orf72 RNA foci as therapy for ALS and frontotemporal degeneration”. Proceedings of the National Academy of Sciences of the United States of America 110 (47), E4530–9. o. DOI:10.1073/pnas.1318835110. PMID 24170860.
- ↑ (2013. október 1.) „Targeting RNA foci in iPSC-derived motor neurons from ALS patients with a C9ORF72 repeat expansion”. Science Translational Medicine 5 (208), 208ra149. o. DOI:10.1126/scitranslmed.3007529. PMID 24154603.
- ↑ (2012. augusztus 1.) „Targeting nuclear RNA for in vivo correction of myotonic dystrophy”. Nature 488 (7409), 111–5. o. DOI:10.1038/nature11362. PMID 22859208.
- ↑ (2012. március 1.) „RNase H-mediated degradation of toxic RNA in myotonic dystrophy type 1”. Proceedings of the National Academy of Sciences of the United States of America 109 (11), 4221–6. o. DOI:10.1073/pnas.1117019109. PMID 22371589.
- ↑ (2011. december 1.) „Potent and selective antisense oligonucleotides targeting single-nucleotide polymorphisms in the Huntington disease gene / allele-specific silencing of mutant huntingtin”. Molecular Therapy 19 (12), 2178–85. o. DOI:10.1038/mt.2011.201. PMID 21971427.
- ↑ (2010. november 1.) „Allele-selective inhibition of mutant huntingtin expression with antisense oligonucleotides targeting the expanded CAG repeat”. Biochemistry 49 (47), 10166–78. o. DOI:10.1021/bi101208k. PMID 21028906.
- ↑ (2009. január 1.) „Identification of a new G-quadruplex motif in the KRAS promoter and design of pyrene-modified G4-decoys with antiproliferative activity in pancreatic cancer cells”. Journal of Medicinal Chemistry 52 (2), 564–568. o. DOI:10.1021/jm800874t. PMID 19099510.
- ↑ (2013. április 1.) „MAZ-binding G4-decoy with locked nucleic acid and twisted intercalating nucleic acid modifications suppresses KRAS in pancreatic cancer cells and delays tumor growth in mice”. Nucleic Acids Research 41 (7), 4049–4064. o. DOI:10.1093/nar/gkt127. PMID 23471001.
- ↑ (2009. március 1.) „Selectivity in ligand recognition of G-quadruplex loops”. Biochemistry 48 (8), 1675–80. o. DOI:10.1021/bi802233v. PMID 19173611.
- ↑ (2014. június 1.) „Small-molecule quadruplex-targeted drug discovery”. Bioorganic & Medicinal Chemistry Letters 24 (12), 2602–12. o. DOI:10.1016/j.bmcl.2014.04.029. PMID 24814531.
- ↑ (2014. február 1.) „TMPyP4 porphyrin distorts RNA G-quadruplex structures of the disease-associated r(GGGGCC)n repeat of the C9orf72 gene and blocks interaction of RNA-binding proteins”. The Journal of Biological Chemistry 289 (8), 4653–9. o. DOI:10.1074/jbc.C113.502336. PMID 24371143.
- ↑ R. Vilar.szerk.: A. Sigel, H. Sigel, E. Freisinger, R. K. Sigel: Chapter 12. Nucleic Acid Quadruplexes and Metallo-Drugs, Metallo-Drugs: Development and Action of Anticancer Agents, 325–349. o.. DOI: 10.1515/9783110470734-018 (2018). ISBN 9783110470734
Források
- (2011. október 1.) „Kinetically grafting G-quadruplexes onto DNA nanostructures for structure and function encoding via a DNA machine”. Chemical Communications 47 (38), 10563–5. o. DOI:10.1039/c1cc13973h. PMID 21858307.
- (2008. augusztus 1.) „In vivo veritas: using yeast to probe the biological functions of G-quadruplexes”. Biochimie 90 (8), 1250–63. o. DOI:10.1016/j.biochi.2008.02.013. PMID 18331848.
- J. L. Huppert, S. Balasubramanian (2005). „Prevalence of quadruplexes in the human genome”. Nucleic Acids Research 33 (9), 2908–16. o. DOI:10.1093/nar/gki609. PMID 15914667.
- A. K. Todd, M. Johnston, S. Neidle (2005). „Highly prevalent putative quadruplex sequence motifs in human DNA”. Nucleic Acids Research 33 (9), 2901–7. o. DOI:10.1093/nar/gki553. PMID 15914666.
- (2006) „Quadruplex DNA: sequence, topology and structure”. Nucleic Acids Research 34 (19), 5402–15. o. DOI:10.1093/nar/gkl655. PMID 17012276.
- (2002. szeptember 1.) „Direct evidence for a G-quadruplex in a promoter region and its targeting with a small molecule to repress c-MYC transcription”. Proceedings of the National Academy of Sciences of the United States of America 99 (18), 11593–8. o. DOI:10.1073/pnas.182256799. PMID 12195017.
- (2006. május 1.) „Genome-wide prediction of G4 DNA as regulatory motifs: role in Escherichia coli global regulation”. Genome Research 16 (5), 644–55. o. DOI:10.1101/gr.4508806. PMID 16651665.
- (2009. június 1.) „A biomimetic potassium responsive nanochannel: G-quadruplex DNA conformational switching in a synthetic nanopore”. Journal of the American Chemical Society 131 (22), 7800–5. o. DOI:10.1021/ja901574c. PMID 19435350.
- szerk.: Neidle & Balasubramanian: Quadruplex Nucleic Acids. Royal Society of Chemistry (2006). ISBN 978-0-85404-374-3
- (2013. december 1.) „The effect of pyridyl substituents on the thermodynamics of porphyrin binding to G-quadruplex DNA”. Bioorganic & Medicinal Chemistry 21 (23), 7515–22. o. DOI:10.1016/j.bmc.2013.09.036. PMID 24148836.
További információk
- Nanopore and Aptamer Biosensor group{NAB group}
- G-Quadruplex World – a G-kvadruplexekről szóló tanulmányok és más információk megbeszéléséről szóló lap
- Greglist – G-kvadruplex-szabályozott gének adatbázisa
- Database on Quadruplex information: QuadBase from IGIB
- GRSDB – RNS-feldolgozó helyekhez közeli G-kvadruplexek adatbázisa.
- GRS_UTRdb Archiválva 2011. július 20-i dátummal a Wayback Machine-ben. - UTR-beli G-kvadruplexek adatbázisa.
- G-quadruplex Resource Site
- non-B Motif Search Tool at non-B DB – G-kvadruplex- és más nem B-DNS-alkotó szekvenciák előrejelzése DNS-szekvenciákból
- QGRS Mapper: a web-based application for predicting G-quadruplexes in nucleotide sequences and NCBI genes.
- Quadfinder: Tool for Prediction and Analysis of G Quadruplex Motifs in DNA/RNA Sequences from Maiti's group, IGIB, Delhi, India[halott link]
- G4Hunter
- pqsfinder
- pqsfinder: onilne kereső a legfrissebb R/Bioconductor-csomaggal
